Caio Fonseca
Computer Science PhD Student
Waterford Institute of Technology
Walter Institute for Information and Communication Systems Science
Biography
Caio Fonseca is a PhD Student at the Walton Institute for Information and Communication Systems Science, Waterford Institute of Technology - WIT, working within the EU-H2020-FET GLADIATOR research project. His PhD work is supervised by Dr. Sasi Balasubramaniam and Dr. Michael Barros.
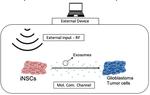
Caio’s research is currently related to mathematical and computational modeling of molecular communications systems in the brain for the treatment of brain pathologies, especifically, glioblastoma multiform. Also to develop evolutionary algorithms for the programming and design of brain organoids.
He is an Electrical and Electronics Engineer and a Computational Scientist. He received his Bachelor’s degree in Electrical Engineering with a concentration in Power Engineering from the Federal University of Campina Grande - UFCG in 2019.
In his undergraduate studies he was an exchange student with a scholarship from the Brazilian Scientific Mobility Program, studying at the Western New England University, Springfield, MA - USA in 2014 - 2015. He was a Visiting Undergraduate Researcher at the University of California at Los Angeles - UCLA in 2015. In 2017, he was also a Visiting Researcher at Edmond & Lily Safra International Institute of Neurosciences working within a group of engineers and physicians.
- Molecular Communications & Information Theory
- Neural and Bioengineering
- Molecular Biophysics/Nanobioscience
- Computational Biology/Neuroscience
- Bio-inspired/Evolutionary Computing
- Cancer Targeted Drug-Delivery
- Nanomedicine/Precision Medicine
- Artificial Intelligence
- Genetic Engineering/Bioinformatics
- Electrical and Electronics Engineering/Power Electronics
- Control Engineering
PhD in Computer Science, 2023
Waterford Institute of Technology - WIT, Ireland
BEng in Electrical Engineering, 2019
Federal University of Campina Grande - UFCG, Brazil
Exchange Program in Electrical Engineering, 2014-2015
Western New England University - WNE, USA